BluePippin Bulletin: Size Selection of 40kb and 50kb DNA
Well, we’ve finally gone where no automated DNA size selection provider has gone before. Using compression-busting pulse-field programs and fancy gel formulations, we’ve tackled the large cut.
Our latest cassette type, BHF7510, comes with four cassette definitions:
1. High Range 40kb Marker W1- Tight
2. High Range 40kb Marker W1- Broad
3. High Range 50kb Marker W1- Tight
4. High Range 50kb Marker W1- Broad
In all four cases, the size selection is centered on the 40kb or 50kb target; range mode selections are not allowed. The “Tight” definitions provide very narrow distributions, while the “Broad” definition doubles the duration of elution to provide higher yields. Below is a summary of some our validation data.
Thanks to our crack R&D team for unlocking the mysteries of HMW fragments!
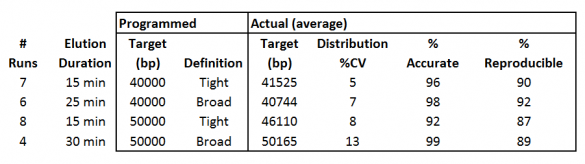
Figure 1. Data summary from High-Range cassette validation tests. Eluted DNA was run on agarose gels using the Pippin Pulse system (3-70kb preset program) and analyzed with the TL100 analysis software from Totallab, Ltd.
From the Trenches: PacBio User Shares Experience with BluePippin
As the BluePippin makes its way to more users of the Pacific Biosciences sequencing platform, there is great data emerging on how our high-pass size selection protocol really complements the sequencer.
A particularly nice example of this comes from Lex Nederbragt, research fellow at the University of Oslo and member of the Norwegian High-Throughput Sequencing Centre, who recently put up a blog post about his experiences with BluePippin and PacBio. He includes plenty of data showing sequencing runs with and without BluePippin selection.
Nederbragt writes that BluePippin was very effective at reducing small fragments from the library, boosting average read lengths and longest subread lengths, and shifting base distribution significantly to the longest reads. Be sure to check out the figures in his blog post for specific data.
Many thanks to the Norwegian High-Throughput Sequencing Centre for their work with BluePippin and to Nederbragt for sharing this information!
Back from Baltimore and the Stellar PacBio UGM
Earlier this week Pacific Biosciences hosted a user group meeting at the University of Maryland in Baltimore, and we were pleased to participate in and co-sponsor the event. As our blog readers know, we recently announced a co-marketing partnership with PacBio to provide BluePippin size selection to PacBio users to help them generate longer average read lengths.
At the user group meeting, about 100 attendees came to discuss their own experiences with the sequencing platform and to learn how other people were pushing the boundaries on read length, DNA input requirements, and more. It was great to see so much enthusiasm for PacBio’s sequencer. This is a really engaged community — almost every presentation was followed by several questions and often lively discussion within the audience as people shared their own methods or results.
A key theme at the meeting revolved around the dramatic improvements in throughput and read length since the PacBio system first launched. Speakers got in the habit of noting what year certain data had been generated as shorthand to help attendees understand how the data might look today if the experiment were run with newer enzymes, reagents, or hardware. A speaker from Baylor noted that since receiving their instrument, throughput has increased more than 10-fold while average read length has jumped from 1.5 Kb to 7 Kb. Here at Sage, we are proud to be part of that growth in read length — and we were glad to see speakers reporting a significant boost in average read length with the incorporation of BluePippin into the workflow.
With these extraordinarily long reads, many of the speakers presented information on genetic elements or attributes that have never been seen before, including connections of distant tandem repeats, genome-wide methylation in pathogens, and fully sequenced mRNA transcripts. There was also quite a bit of excitement around moving toward de novo mammalian sequencing with the PacBio workflow, which can currently perform de novo sequencing and automated assembly/finishing of microbial genomes.
From Sage Science, Chris Boles offered an update on BluePippin for PacBio. (For the current protocol, click here.) He presented data from one genome center showing a project in which the N50 median read length of 4,800 bases was boosted to 9,100 bases by adding BluePippin to remove smaller fragments from the library. Boles also reported that we are working to develop additional protocols with PacBio to keep extending those N50 numbers. We look forward to continuing to help PacBio users generate even more impressive read lengths from their sequencers!
ASMS, Here We Come!
We are packing our bags for Minneapolis and the 61st annual conference of the American Society for Mass Spectrometry. As scientists who come from the DNA realm, the protein world is very exciting to us and we’re looking forward to one of the premier meetings in this field.
It’ll be great to see the award presentation from Dick Smith, head of proteomics research at Pacific Northwest National Laboratory. Dick has a long history of optimizing mass spec technology, so no doubt his talk will cover his greatest hits as well as what’s new and intriguing in his lab today. We’re also interested in hearing about cutting-edge work in top-down proteomics and other approaches during the regular conference sessions.
This is the first year Sage will have a booth at ASMS, so please stop by booth #31 and say hello! We’ll be introducing our two new automated tools for the protein market: the BluePippin for proteins and the Sage ELF. BluePippin, one of our DNA sizing platforms, has new cassettes to enable targeted SDS-protein collection of a desired fraction from up to five samples per run. The ELF, or Electrophoretic Lateral Fractionator, allows users to fractionate a protein sample into 12 contiguous size fractions. These tools provide increased reproducibility and ease-of-use for procedures that require gel isolation of proteins, including top-down and bottom-up proteomics studies, post-translational modification analysis, and other protein mass spectrometry methods.
You can also check out the two posters we’ll have, both during Wednesday’s poster session.
• Poster #590: “Using Orthogonal Techniques for Protein-Peptide Separation to Generate Comprehensive HDMSe Mass Spectral Libraries from an E. coli Model System”
• Poster #674: “The BluePippin Automated Size-Fractionation System for Proteins”
We hope to see you there!
New Prostate Cancer Research Indicates Rapid Accumulation of DNA Mutations
Researchers studying the behavior of prostate cancer have shed light on how the cancer develops mutations so rapidly. In contrast to other cancers that seem to progress by adding mutations one at a time, the prostate cancer samples that were analyzed showed evidence of a rapid chromosomal scramble resulting in several alterations that disrupt many genes at once. The scientists named this phenomenon “chromoplexy.”
The findings are described in “Punctuated Evolution of Prostate Cancer Genomes,” published in Cell by a large collaboration of scientists with senior authors Francesca Demichelis, Mark Rubin, and Levi Garraway. We were delighted to see that the team used Pippin Prep automated size selection as part of their sample preparation workflow in this impressive research.
For this work, the team sequenced 57 tumor genomes and their matched normal tissue, hoping to build on findings from earlier exome studies and to provide a better understanding of the onset and progression of prostate cancer. They also analyzed copy number changes across the genomes and studied the chromoplexy phenomenon they found, through which large chunks of the genomes had been jumbled in a short period of time. “These complex rearrangement events occur in the majority of prostate cancers and may commonly inactivate multiple tumor-constraining genes in a coordinated fashion,” the authors note.
They add, “Our modeling suggests that chromoplexy may induce considerable genomic derangement over relatively few events in prostate cancer and other neoplasms, supporting a model of punctuated cancer evolution.”





