Pippin and PacBio: Better Together
Update: Just announced– Sage Science and Pacific Biosciences have partnered to offer users longer reads than ever. Click here to read the press release.
The new SMRTbellTM 20kb sample prep protocol using the BluePippin is posted on the PacBio® SampleNet site, click here to view the page or download the document. Instructions for using the BluePippin High Pass cassette definition for the SMRTbell protocol can be found here.
Original Post:
Many thanks to all the scientists who stopped by the Sage suite during the Advances in Genome Biology & Technology meeting in Marco Island — we were thrilled to see you! When we weren’t busy handing out glowing medallions,  the Sage Science team had a great time catching up with people, attending the excellent scientific presentations, and walking the halls of the poster sessions.
the Sage Science team had a great time catching up with people, attending the excellent scientific presentations, and walking the halls of the poster sessions.
Along with other attendees, we were really impressed by the posters and presentations showing data from the PacBio® RS. A number of talks referred to the company’s single-molecule, real-time (SMRT®) sequencing, which generates extremely long reads — one scientist said he’d produced a 26 kb read on the PacBio RS. Between the read length and unique base modification detection capability, the sequencing platform is particularly useful for de novo assembly, genome finishing, SNP discovery, and epigenetic analysis.
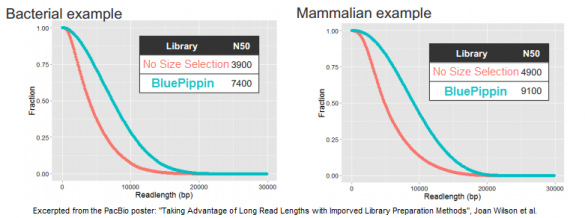
In its quest to produce sequence from even longer DNA fragments, PacBio recently tested our BluePippin platform for size selection; the results of those studies were the basis for a poster we displayed at AGBT. In it, the PacBio authors demonstrate a doubling of the N50 subread length – the value at which 50% of unique DNA bases are in reads longer than this value – when using the BluePippin platform compared to the standard workflow (click to enlarge this excerpt from the poster):
For generating 20 kb libraries that are optimal for the platform’s current polymerase, the authors note that “when combined with size selection to remove the shorter fragments that tend to load more favorably than long fragments, these libraries show greatly increased subread lengths.”
Looking at three examples of libraries for bacterial, fish, and mammalian DNA, the authors report that the N50 subread lengths increased by 90%, 110%, and 86%, respectively. Some of this data was also included in an AGBT plenary presentation by Jonas Korlach, Chief Scientific Officer of PacBio, who cited the BluePippin platform as one of the new improvements for PacBio customers.
Some PacBio customers are already using the BluePippin platform, and we look forward to working with more of them in the coming months. Together, these technologies offer a great and unique solution for truly long-read sequencing.
Pippin Pulse
Field Inversion Gel Electrophoresis
We’ve set up this post to provide Pippin Pulse users with a resource to keep updated on new protocols and to post references that might be useful. We also invite users to feel use this page as a forum, if you see fit, and we’d enjoy showing interesting gel images and data (email these to support@sagescience.com).
Our go-to book on theory:
“Electrophoresis of Large DNA Molecules:Theory and Applications” (E. Lai and B. Birren, eds), Current Communication in Cell & Molecular Biology Vol. 1,. Cold Spring Harbor Laboratory Press, New York, 1990
Download the Pippin Pulse user manual here
Our Gel Set-up
We use a Galileo Model 1214 RapidCastTM Mini Gel Unit running 12 X 14 cm gels using Lonza SeaKem® Gold Agarose. We use our Pippin Tris-TAPS buffers (which you can order directly from us in the US and Canada, Part No. KBB1001, under “Accessories” on our ordering page) or 0.5X TBE. The formulations can be found in the Pippin Pulse user manual or separately here.
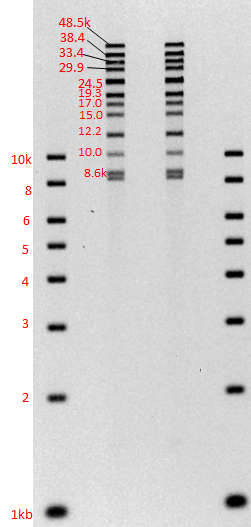
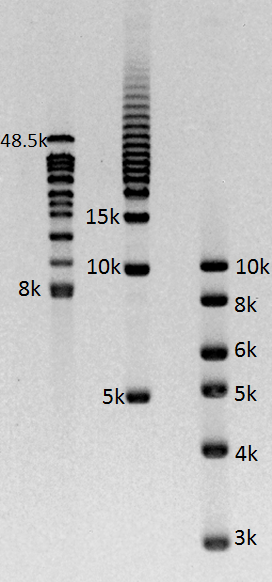
Gel images of our 1-50kb and 3-70kb protocols:
(click to enlarge images)
Improved Recovery for 3-10kb DNA Size Selection
BluePippin users, we have a new cassette definition that will appear in the Cassette Definition Set 10 (CD10) that we feel merits some further detail. This definition is named “0.75% DF 3 – 10kb Marker S1 – Improved Recovery”
What is it?
This definition improves the sample recovery for customers using BLF7510 Marker S1 cassette kits for DNA size selection, for size ranges between 3- 10 kb. The original cassette definition for this range, “0.75% DF 3 – 10kb Marker S1” uses pulsed-field during elution (size selection), and we have subsequently found that direct-current elution improves recovery.
Who should use it?
This is now the recommended cassette definition to use for size-selecting between 3-10kb. However, if you have optimized a protocol using the current version you may want continue to use it (it will not be deleted as a cassette definition option) if you do not wish to re-optimize.
Also note that since the BluePippin system does not allow for simultaneous pulsed-field and direct-current operation, lanes that are being separated using pulsed-field will temporarily be halted during every elution process. This makes the total run times lengths dependent on the elution times (and size selection targets), and therefore more difficult to estimate and can add up to 40 minutes to a cassette run.
How much improved is the recovery?
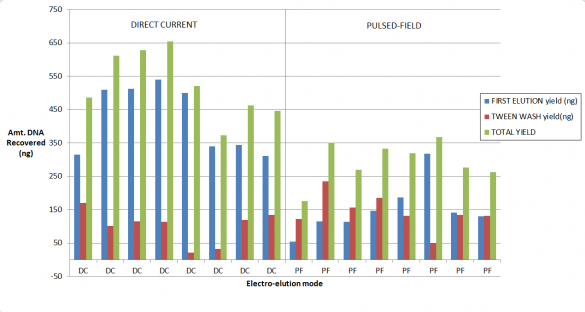
Our validation of the change in the cassette definition shows up to a 2-fold increase in recovery. As you may know, for collections of large fragments (on 0.75% agarose cassettes) we provide Tween solution which we find improves recovery if one employs a post-collection Tween rinse. Considered together, here are some results from our study. We started with a 5 ug of restriction digested DNA, and 8kb “Tight” collections:
At TGAC, Pippin-Aided Sequencing Gives Quality Boost to Genome Assemblies
Before a new technology is deployed at The Genome Analysis Centre in Norwich, UK, it must run the gauntlet of Matthew Clark’s lab.
Clark is the sequencing technology development leader at TGAC, where part of his role involves testing out new platforms, deciding whether they’d be a good fit at the institute, and ironing out best-practice workflows for the ones that are chosen.
The BluePippin automated size selection platform from Sage Science is one of the tools that succeeded in Clark’s technology proving ground and is now used more broadly throughout TGAC, where major projects include sequencing the wheat genome and studying honeybees. Clark, who joined the institute in 2010 after spending seven years as a scientist at the Wellcome Trust Sanger Institute, says that TGAC researchers rely on Pippin for building long-insert libraries for sequencing projects.
These projects include de novo sequencing, for which precise size selection in mate-pair libraries offers a significant improvement in quality of the genome assembly. The mate-pair sequence might represent a small fraction of the total data generated for the assembly — most will come from the shorter-insert paired-end libraries — “but it has a massive effect on the quality of the output,” Clark says. “The bigger-insert library gives you a 5x or 10x jump in quality, maybe even bigger, in terms of the sizes of the assembly that you’re able to generate.” These libraries, which are sequenced on Illumina or Pacific Biosciences instruments, offer longer-range information that can fill gaps or jump over repeats and other problematic structures that would otherwise break up an assembly. Indeed, Clark says the TGAC bioinformatics team prefers Pippin-aided sequencing libraries because the tight size selection helps them determine how far apart certain reads should be and put together a more accurate assembly.
Clark’s team had tested other automated size selection options, but Pippin was the best platform for generating these valuable large-insert libraries. (Pippin Prep can yield libraries with inserts up to 8 kb, while BluePippin works for even longer inserts.) It also allows for loading more material than other size selection alternatives, making it a good fit for the no-PCR paired-end libraries that some TGAC scientists like to run, Clark says. “If you have enough DNA, you can just skip the PCR — and you get much better coverage across the genome and a better assembly. That’s easier to do on a Pippin.”
NHGRI Scientists Find Transgenic DNA Has More Rearrangements than Expected
Amanda DuBose didn’t set out to come up with a new technical solution to characterizing transgenic DNA in animal models. A postdoctoral fellow in Francis Collins’ lab at the National Human Genome Research Institute, DuBose was using mice to study the premature aging disease Hutchinson-Gilford Progeria Syndrome (HGPS) when she and her colleagues realized that they were hindered by their inability to determine which mice were homozygous for the transgene containing the mutated gene of interest. This was crucial information: mice that are homozygous for the transgene with the HGPS-causing variant have a more severe phenotype with an earlier onset, making them more useful for research than the heterozygotes.
The scientists tried a number of tools — including qPCR, FISH, and Southern blots — but none of the assays could quickly and reliably report the number of gene copies in a mouse. So the team resorted to what DuBose calls “an invention of necessity” and made a custom microarray to identify the site of transgene integration and develop a genotyping assay that could deliver the results they needed.
The results of that work were published in Nucleic Acids Research in a paper entitled “Use of microarray hybrid capture and next-generation sequencing to identify the anatomy of a transgene.” Lead author DuBose and her colleagues analyzed regions of the transgenic mouse genome next to BAC sequence by using arrays to capture the DNA, size selecting with the Pippin Prep, and sequencing on the Illumina HiSeq.
DuBose, Collins, and the other authors found that the BAC had broken more than expected when it was injected into the mouse; there were four mouse-human junctions. “I was surprised by how rearranged it was,” DuBose says, noting that the junctions within the BAC were quite unexpected. “We know it’s not unique to our model. I have a feeling that all of the transgenic models have crazy rearrangements — it’s just that nobody goes in to look this closely.”
Ultimately, DuBose used the sequence data to design a PCR assay with three primers crossing one of the BAC-mouse junctions. That assay is now a standard tool that can be used to determine gene copy number in the model mice: results indicate whether a mouse is homozygous mutant, heterozygous mutant, or wild type.
The lab has now returned to its HGPS research, but DuBose says that the technological approach to characterizing a transgene and developing a PCR genotyping assay can be used by anyone working on transgenic models or genome sequence that has foreign DNA in it. She notes that the array her team used for targeted capture is no longer available, so the technique would have to be adapted for liquid capture.