PacBio and Sage Science
HiFi sequencing has set the bar for gold-standard whole genome analysis. Consensus long reads provide highly accurate sequences and sufficient read length to capture phasing information as well as structural variants and repetitive elements. (Preparing whole genome and metagenome libraries using SMRTbell® prep kit 3.0, Procedure and checklist)
Sage Science’s electrophoretic size selection approach can be a key factor in achieving the best results possible. It is particularly adept at rejecting small DNA from a library and thereby allowing optimally sized fragments —based on the processivity of the polymerase— to be read by the zero mode waveguides.
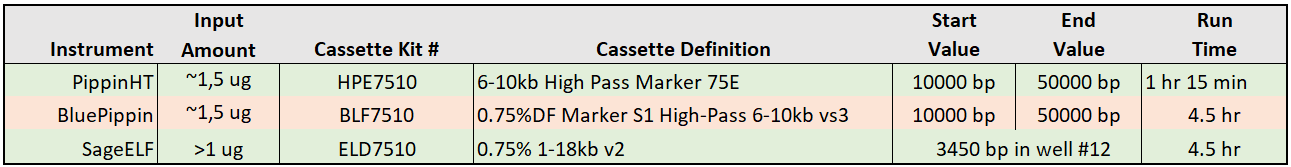
PacBio provides size selection parameters for three Sage instruments in their Alternative Size Selection Methods For SMRTbell® Prep Kit 3.0. These are summarized in the Table below:
Which Size Selection instrument to choose?

The PippinHT system is the top choice for higher-throughput workflows. The PippinHT can process 11 size selections at once on a gel cassette (1 of the 12 lanes must be dedicated to an external size marker). The PippinHT accommodates 2 gel cassettes so up to 22 samples can be run at once, and at a quarter of the run time (1 hr 15 m) when compared to the BluePippin or SagELF. Less than 11 samples can be run on a single cassette as well, provided that the cassette is resealed (tightly) with PCR tape after use. If doing this, it is important to remember to turn off unused lanes in software during the initial run and that a marker must again be used with the re-run.
 The BluePippin can process 4 size selections per run, with a fifth lane dedicated to an external size marker. One key feature to note is that the BluePippin will accept DNA sample inputs up to 5 ug (vs. 1.5 ug for the PippinHT), which some users may wish to exploit.
The BluePippin can process 4 size selections per run, with a fifth lane dedicated to an external size marker. One key feature to note is that the BluePippin will accept DNA sample inputs up to 5 ug (vs. 1.5 ug for the PippinHT), which some users may wish to exploit.
 The SageELF instrument fractionates a single DNA Sample into 12 contiguous size bins and runs a single sample per cassette and up to 2 cassettes at once. HiFi users appreciate its ability to achieve relatively narrower fragment size distributions than the Pippin line. To use the system, users must analyze the contents of the size bins since the sizes are estimated in software and may not be precise. This actually provides some additional flexibility as size ranges can be built by pooling adjacent lanes, and unused size bins can be saved and stored for later analysis.
The SageELF instrument fractionates a single DNA Sample into 12 contiguous size bins and runs a single sample per cassette and up to 2 cassettes at once. HiFi users appreciate its ability to achieve relatively narrower fragment size distributions than the Pippin line. To use the system, users must analyze the contents of the size bins since the sizes are estimated in software and may not be precise. This actually provides some additional flexibility as size ranges can be built by pooling adjacent lanes, and unused size bins can be saved and stored for later analysis.
Resources
Technical Support?
Don’t hesitate to contact us! 888-744-0144 or email us: support@sagescience.com
Check out our most common tech support FAQs
Latest App Notes
Higher Accuracy “Range + T” Size Selection: BluePippin and PippinHT
Recent HiFi Citations featuring Sage Science Products
Check them out here.






