Many thanks to all the scientists who stopped by the Sage suite during the Advances in Genome Biology & Technology meeting in Marco Island — we were thrilled to see you! When we weren’t busy handing out glowing medallions,  the Sage Science team had a great time catching up with people, attending the excellent scientific presentations, and walking the halls of the poster sessions.
the Sage Science team had a great time catching up with people, attending the excellent scientific presentations, and walking the halls of the poster sessions.
Along with other attendees, we were really impressed by the posters and presentations showing data from the Pacific Biosciences sequencer. A number of talks referred to the company’s single-molecule, real-time (SMRT®) sequencing, which generates extremely long reads — one scientist said he’d produced a 26 Kb read on the PacBio RS. Between the read length and unique base modification detection capability, the sequencing platform is particularly useful for de novo assembly, genome finishing, SNP discovery, and epigenetic analysis.
In its quest to produce even longer reads, PacBio recently tested our BluePippin platform for size selection; the results of those studies were the basis for a poster we displayed at AGBT. In it, the PacBio authors demonstrate a doubling of median read length when using Pippin compared to the standard workflow (click to enlarge this excerpt from the poster):
For generating 20 Kb libraries that are optimal for the sequencer’s new XL polymerase, the authors note that “when combined with size selection to remove the shorter fragments that tend to load more favorably than long fragments, these libraries show greatly increased subread lengths.”
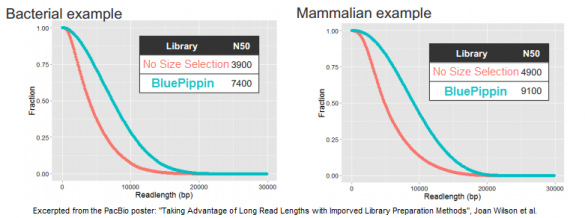
Looking at three examples of libraries for bacterial, fish, and mammalian DNA, the authors report that N50 read lengths — the halfway point between shorter and longer reads — increased by 90%, 110%, and 86%, respectively. Some of this data was also included in an AGBT plenary presentation by Jonas Korlach, Chief Scientific Officer of PacBio, who cited BluePippin as one of the new improvements for PacBio customers.
Some PacBio customers are already using the Pippin platform, and we look forward to working with more of them in the coming months. Together, these technologies offer a great and unique solution for truly long-read sequencing.